Pu·MA System 3D for Automated Organoid Assays, In Situ Sampling and In Situ Imaging
Introduction
3D cell models and organoids provide a better representation of in vivo tissue or organ function. For the last three decades, researchers have been perfecting the formation and maintenance of various 3D models for understanding both disease and normal physiology (1-3). Some of the limiting factors have been the ability to perform complex assays easily and quickly with these precious samples especially patient-derived material. Also, when manually performing drug treat-ments and assays in 96-well plates, one is limited by the number of readouts per sample. The manual treatment, staining, and processing of spheroids and organoids is typically labor-intensive and prone to disrup tion or loss of samples.
In this application note, we report the use of our microfluidic-based Pu·MA® System to perform automated assays using 3D cell models. The spheroids in this study HeLa (cervical carcinoma line) and HepG2 (hepatocyte carcinoma line) were incubated with and without com-pounds for 24−48 hours in the Pu·MA System. After drug treatments, the spheroid media was collected for secreted factor analysis. The spheroids were then stained and imaged within the flowchips using a confocal microscope. Other cell viability measurements or downstream analysis can be performed.
Pu·MA System Workflow
Pu·MA System and flowchips have been designed for a streamlined workflow as shown in in Figure 1. This consists of:
- loading spheroids and reagents into the flow-chips
- placing the holder into the Pu·MA System (in-side the incubator)
- running assay protocols via an intuitive touch-screen interface
- analyzing the spheroids within the flowchip us-ing a confocal microscope
- sampling supernatant for downstream analysis
The system architecture and use of pneumatics to move fluids provides gas exchange to the sample chambers within the incubator environment.
The flowchip features include:
- a convenient multi-well plate format (384-well spacings SLAS/ANSI standard)
- standard spacing for use with multichannel pipettes or automated liquid dispense system
- up to 32 tests per plate
- optically clear bottoms for imaging with any fluorescence or confocal imaging system
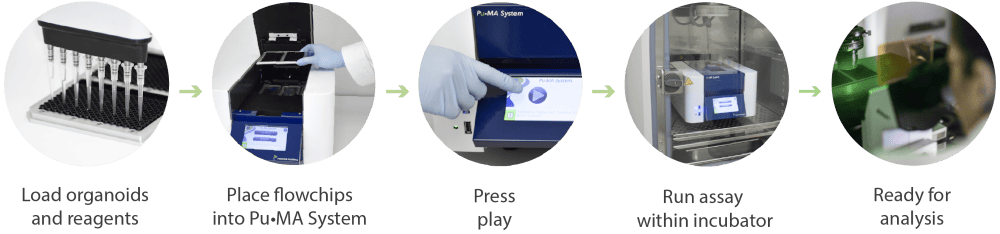
Figure 1. Schematic of the Pu·MA System workflow for automated organoid assays, in situ staining and imaging.
