Use of Pu·MA System 3D for Single Spheroid Assays with Downstream Metabolomics
Introduction
There is an increasing interest in using three-dimensional (3D) cell structures for modeling tumors, organs, and tissue to accelerate translation research1. Significant progress has been made in formation of such structures to recapitulate the in vivo environment but performing complex assays with them can be challenging. For example, manual treatment, staining, and processing of spheroids and organoids is typically labor-intensive and prone to disruption or loss of samples. Here we report on use of our microfluidic-based Pu·MA® System to perform automated assays with single spheroids followed by metabolite sampling using a novel in situ lysing technique. Metabolomic profiles of spheroids with and without treatment of compounds were obtained showing significant differences in their profiles.
Pu·MA System Workflow
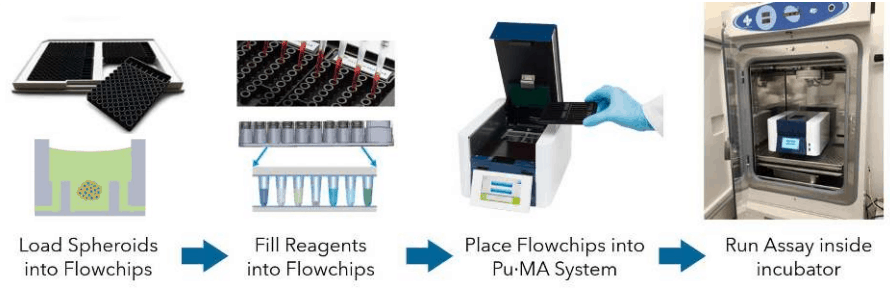
Pu·MA System Flowchips are designed with chambers and reservoirs arranged in a convenient multi-well plate format (384-well spacings) and provide up to 32 tests per plate. Once spheroids and reagents are loaded into the flowchips, the plate is placed into the Pu·MA System and reagent exchanges are done automatically through microfluidic channels connected to the protected sample chamber (Fig 1). Multiple reagent exchanges can be performed enabling complex assay protocols to be run.
Assay programs are pre-loaded into the system and run using an intuitive touch-screen interface. The whole Pu·MA System can be placed in an incubator to run assays at 37°C and 5% CO2. The system architecture and use of pneumatics to move fluids provides gas exchange to the sample chambers.
Figure 1. Schematic of the Pu·MA System workflow.